The Clinical Data Accuracy Gap: Why Secondary Use of EHR Data Produces Wrong Results
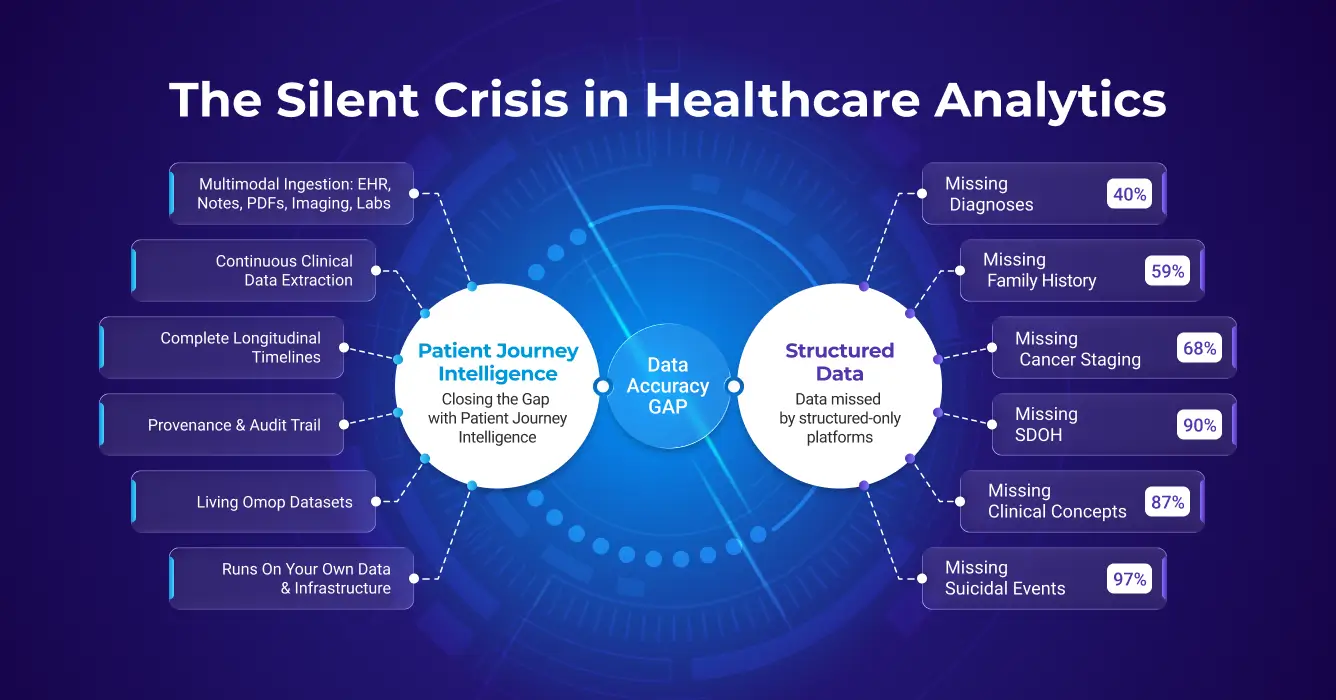
TL;DR: The Clinical Data Accuracy Gap refers to the 40%-87% of patient information, including diagnoses, SDOH, and cancer staging, that is documented in unstructured clinical notes but missing from structured EHR fields, leading to biased analytics and underpowered AI models.
The clinical data accuracy gap is the systematic inaccuracy that emerges when healthcare analytics and AI projects operate on incomplete patient data. Secondary use, which means repurposing clinical data collected during care for research, registries, population health, and AI model development, depends entirely on whether that data accurately represents what actually happened to patients. Across dozens of peer-reviewed studies, the evidence is consistent: structured EHR data alone misses 40–87% of clinically relevant information, producing cohorts that are biased, models that underperform, and results that can mislead clinical and regulatory decisions.
Key Statistics at a Glance: Structured EHR vs. Unstructured Clinical Notes
| Clinical Data Type | Structured EHR Capture | Unstructured Notes | Invisible to Structured Queries | Clinical Implication |
|---|---|---|---|---|
| Diagnoses | ~60% | ~40% found only in notes | 40% missed | Cohort queries miss nearly half of eligible patients; clinical trial recruitment and population health programs (utilizing ICD-10 and SNOMED standards) are systematically underpowered. |
| Family History | ~5% | ~59% | 12x discrepancy | Risk stratification models built on structured data alone lose their most heritable predictive signal, producing unreliable genetic risk scores. |
| SDOH (Social Determinants) | 2% (ICD-10 Z-codes) | 93.8% (NLP on notes) | 46x discrepancy | Readmission and outcome models ignore the social drivers that explain most variation, making interventions ineffective and equity analyses impossible. |
| Cancer Staging | <32% | >68% missing from structured fields | 68%+ gap | Cancer registries and real-world evidence studies cannot accurately report stage distribution or treatment outcomes without pathology narrative extraction. |
| Medication Histories | 30–40% accurate | 60–70% contain errors | Majority inaccurate | Pharmacovigilance studies, adherence analyses, and drug safety signals derived from structured medication records are built on a foundation that is wrong more often than right. |
| Suicide / Self-harm Events | <19% coded | >81% only in notes | >81% missed | Mental health risk models and quality measures relying on ICD codes miss the overwhelming majority of at-risk patients, with direct patient safety consequences. |
| All Clinical Concepts | 13% have structured counterparts | 87% in free text only | 87% gap | Any AI model trained on structured EHR fields alone operates on less than one-seventh of the available clinical signal, making high accuracy on complex clinical tasks structurally unachievable. |
Sources: Poulos et al. 2021, Polubriaginof et al. 2015, Guevara et al. 2024, Emamekhoo et al. 2022, Fernandes et al. 2018, Seinen et al. 2025
What makes secondary use hard is that structured clinical data (e.g. EHR, claims) was never designed for secondary use. It was captured to support billing, documentation, and care coordination, not analytics or AI. The result is a dataset that is structurally incomplete by design.
Unlike operational systems designed to support real-time clinical workflows, prescribing medications, ordering labs, documenting visits, secondary use environments must work with years or decades of historically accumulated data. This data comes from multiple systems, uses inconsistent formats and terminologies, includes both structured and unstructured content, and often contains gaps, conflicts, and quality issues that went unnoticed during original data entry.
The promise of secondary use is compelling: unlock insights hidden in millions of patient encounters, identify at-risk populations before problems escalate, accelerate clinical trial recruitment, automate registry reporting, and train AI models that improve care. But realizing this promise depends entirely on whether the data accurately represents what actually happened to patients, and that's where most secondary use initiatives struggle.
The Clinical Data Accuracy Challenge: Incomplete Patient Views Lead to Wrong Results
Beyond engineering cost, the most serious and pervasive problem with today's secondary use approaches is accuracy. When projects operate on an incomplete view of patient data, they don't merely return partial answers, they often return wrong or misleading results.
Incomplete Patient Data Creates Systematic Inaccuracy
Most healthcare organizations have spent decades accumulating clinical data across dozens of systems: electronic health records, laboratory information systems, radiology platforms, specialty registries, billing systems, scanned document repositories, and more. But integrating all of this data into a unified, longitudinal patient view is expensive and technically complex.
As a result, many secondary use initiatives cannot afford to reconstruct a complete patient journey across all available data modalities. The complexity and cost of integrating free-text notes, scanned documents, imaging metadata, specialty systems, and longitudinal context lead teams to limit scope to what is easiest to access, most commonly, structured EHR data alone.
⚠️ The Structured-Only Trap
Projects often systematically underestimate how much accuracy is lost by relying on a single modality. Structured EHR fields capture only a subset of clinically relevant information and frequently omit context such as certainty, negation, temporality, disease progression, adverse events, social factors, and clinician reasoning.
As a result, analytics, registries, cohorts, and AI models built on structured-only data routinely miss critical signals, misclassify patients, and produce biased or incomplete outputs.
The Research Evidence: How Much Clinical Information Do Structured-Only Systems Miss?
Multiple peer-reviewed, published studies quantify the magnitude of accuracy loss when unstructured and longitudinal data are excluded from secondary use projects:
Clinical Diagnoses: The 40% Gap
~40%
Nearly 40% of important inpatient diagnoses are mentioned only in free-text clinical notes and are absent from structured problem lists (Poulos et al., 2021). If your cohort query searches only structured diagnosis codes, you're systematically missing nearly half of the patients who actually have the condition you're studying.
Family History: The 12x Discrepancy
12×
Family history is documented in unstructured notes for ~59% of patients, but appears in structured fields for only ~5%. That's a 12-fold difference between what clinicians document and what gets captured in structured data (Polubriaginof et al., AMIA Annu Symp Proc. 2015). Any genetics study, risk model, or preventive care program relying on structured family history alone operates with systematically incomplete inputs.
Social Determinants of Health: 93.8% vs. Minority
46×
NLP on clinical notes identified 93.8% of patients with adverse social determinants of health. ICD-10 Z-codes identified just 2.0% of the same patients, which represents a 46-fold difference in detection capability (Guevara et al., npj Digital Medicine. 2024). Housing insecurity, food insecurity, and transportation barriers are documented extensively in clinical notes but almost never coded into structured fields. For any analytics or AI application that relies on SDOH data, structured-only approaches are functionally blind to nearly the entire signal.
Oncology: The Structured Data Gap
In oncology, the richest clinical information, tumor histology, staging details, biomarker results, treatment response, and progression events, exists predominantly in unstructured pathology reports and clinical notes rather than in structured EHR fields (Castellanos et al., JCO Clin Cancer Inform. 2024). This creates systematic gaps in cancer registries, research cohorts, and quality reporting that rely on structured data alone.
>68%
Different studies report that cancer staging data is missing from structured EHR fields in 68%+ of encounters (Emamekhoo et al., JCO Clin Cancer Inform. 2022). TNM staging information is routinely documented in pathology reports and oncology notes but not transferred to structured problem lists or staging fields, making it invisible to automated cohort identification and registry reporting.
>98%
Biomarker data has a median of 14% missing and a mean of 22% missing across molecular epidemiology studies, with some studies reporting up to 98% missing data for key biomarker variables (Desai et al., Cancer Epidemiol Biomarkers Prev. 2011). Critical genomic results (ER/PR/HER2 status, PD-L1 expression, gene mutations) are documented in pathology narratives but frequently absent from structured fields, undermining precision oncology initiatives and outcomes research.
Medication Reconciliation: Discrepancies Everywhere
60% - 70%
Medication reconciliation studies consistently find an average of 60% to 70% of medication histories contain at least one error (Lombardi et al., Rev Lat Am Enfermagem 2016), with over 90% of patients having at least one discrepancy between structured medication lists and what is documented in clinical notes and discharge summaries (Ahmadi et al., Pharmacy. 2024). Structured medication tables alone frequently misrepresent true medication exposure, critical for pharmacovigilance, drug-drug interaction detection, and outcomes research.
Suicidality and Self-Harm: The Coding Gap
>81%
Only 3% of patients with suicidal ideation and 19% with suicide attempts documented in clinical notes have corresponding ICD codes (Fernandes et al., Sci Rep. 2018). This means structured coding systems capture fewer than 1 in 5 documented suicide-related events. Psychiatric screening results, safety plans, and clinician assessments are documented extensively in notes but rarely coded, leaving structured-only surveillance systems blind to the vast majority of high-risk patients.
Clinical Prediction: The Unstructured Data Advantage
87%
Only 13% of clinical concepts extracted from patient records have matching structured counterparts (Seinen et al., J Med Internet Res. 2025). In other words, 87% of extracted clinical information exists solely in free-text narratives, invisible to analytics that query only structured fields. This enormous gap explains why models trained on structured data alone routinely underperform.
Why Text Outperforms Codes: Structured diagnosis codes are designed for billing and administrative classification, not clinical nuance. A patient coded with "diabetes" tells you nothing about glycemic control, medication adherence, complications, or disease trajectory. Clinical notes document "HbA1c 11.2%, poorly controlled despite max dose metformin, recurrent DKA admissions, non-adherent to insulin." This narrative contains the actual predictive signals for outcomes, but remains inaccessible if you query only structured fields.
The Root Cause of Clinical Data Accuracy Gap: Lack of Continuous Multimodal Data Integration
These findings point to a consistent pattern: the dominant failure mode in secondary use projects is not tooling but incomplete patient representation.
Unstructured EHR extraction is the missing step. Without it, the richest layer of clinical documentation - physician notes, discharge summaries, pathology reports, and operative records — remains invisible to analytics and AI. Structured fields capture the administrative skeleton of a patient encounter. Unstructured EHR extraction recovers the clinical substance: the diagnoses reasoned through, the treatments considered and rejected, the progression documented visit by visit.
Secondary use analytics and AI that skip unstructured EHR extraction operate on a partial view of every patient. The result is not just missing data — it is systematically distorted data that biases cohorts, degrades model performance, and produces conclusions that cannot be trusted for clinical or regulatory decisions.
✅ The Path Forward
A modern secondary use data platform must do more than move data, it must fundamentally change how organizations prepare, trust, and reuse clinical data over time. The opportunity is to replace fragmented, per-project pipelines with a single, shared foundation that transforms raw multimodal data into AI-ready patient journeys and keeps them continuously up-to-date as new data arrives.
What a Secondary Use Clinical Data Platform Requires
To achieve accurate secondary use of clinical data, organizations need eight foundational capabilities that go far beyond traditional data warehousing:
1. Multimodal Clinical Data Integration
Requirement: Ingest data across all modalities, structured fields, clinical notes, scanned documents, imaging metadata, labs, medications, procedures, claims.
Why it matters: Relying on any single modality systematically excludes critical information, leading to incomplete cohorts and biased models.
2. Healthcare-Specific NLP
Requirement: Perform NLP medical entity extraction for identifying diagnoses, medications, findings, procedures, and clinical attributes from free-text notes using medical language models, relation extraction, assertion detection, and temporal reasoning.
Why it matters: Healthcare-specific models achieve 85–95% precision/recall and are ~30% more accurate than general-purpose LLMs on clinical tasks.
3. Terminology Standardization
Requirement: Map all clinical concepts to standard medical vocabularies (SNOMED CT, RxNorm, LOINC, ICD-10-CM).
Why it matters: Without normalization, the same clinical fact appears in dozens of forms, making accurate patient counts and cross-institution research impossible.
4. Clinical Reasoning & Conflict Resolution
Requirement: Resolve conflicts, deduplicate entities, distinguish assertion status (confirmed vs. ruled out), reason about temporal changes.
Why it matters: Real-world data is messy, without intelligent reasoning, systems either lose information or create duplicates and noise.
5. Longitudinal Patient Timelines
Requirement: Organize all clinical events chronologically with precise temporal context to support queries about disease progression, treatment response, outcomes.
Why it matters: Most clinical questions involve time, systems need to answer "when" questions, not just "does patient have X?"
6. Privacy & De-Identification
Requirement: Automatically remove PHI from clinical text, documents, and images using HIPAA-compliant methods with 99%+ accuracy.
Why it matters: Most secondary use requires de-identification, manual approaches are slow, expensive, and error-prone. Medical-specific patterns require specialized tools.
7. Provenance & Auditability
Requirement: Track complete lineage from source document to final output. Provide confidence scores. Enable drill-down to source evidence.
Why it matters: Regulatory compliance, research reproducibility, and clinical trust require transparency, not black boxes.
8. Continuous Updates & Living Datasets
Requirement: Keep patient journeys continuously up-to-date as new data arrives, not static snapshots that become stale.
Why it matters: Clinical data accumulates continuously. Quarterly refreshes miss opportunities and make results obsolete.
How Patient Journey Intelligence Solves the Clinical Data Accuracy Gap
The evidence is clear: structured-only approaches systematically miss critical clinical information. But recognizing the problem is different from solving it. Building the infrastructure to ingest multimodal data, extract clinical facts from unstructured text, normalize terminologies, reason about conflicts, and maintain living datasets requires capabilities that most organizations don't have in-house, and shouldn't need to build from scratch.
Patient Journey Intelligence was designed specifically to close this gap. Rather than asking every research team, registry program, or AI initiative to solve the same data accuracy problems independently, the platform provides a shared foundation that transforms fragmented clinical data into complete, trustworthy patient journeys.
Here's how the platform delivers on each of the eight requirements outlined above:
Multimodal Clinical Data Integration
Ingests structured EHR fields, clinical notes, scanned PDFs, imaging metadata, FHIR resources, and claims data through a unified pipeline.
NLP Medical Entity Extraction
Medical Language Models extract diagnoses, medications, and findings with 85–95% precision, understanding negation, uncertainty, and assertion status.
Terminology Standardization
All concepts normalized to SNOMED CT, RxNorm, LOINC, and ICD-10-CM, enabling accurate patient counts and cross-institutional research.
Clinical Reasoning
ML-based confidence scoring, entity deduplication, conflict resolution, and distinction between confirmed diagnoses vs. ruled-out conditions.
Longitudinal Timelines
Clinical events organized chronologically with temporal context for disease progression, treatment response, and time-to-event outcomes.
Privacy & De-Identification
99%+ accurate PHI removal in line with HIPAA and GDPR regulations, configurable for research and external data sharing.
Provenance & Auditability
Complete lineage from source document to OMOP representation with confidence scores and drill-down to supporting evidence.
Continuous Updates
Patient journeys update automatically as new data arrives, ensuring AI agents and analytics operate on current, complete representations.
The result: instead of building incomplete pipelines that miss 40% of diagnoses, 81% of suicidality mentions, and 68% of cancer staging data, organizations build once on a foundation that captures what actually happened to patients, across all modalities, normalized to standard vocabularies, with full provenance and continuous updates.
What You Gain: Accuracy, Speed, and Reuse
By deploying a platform that meets these requirements, organizations unlock transformative improvements in secondary use:
Speed to Insight
Patient timeline construction that previously took weeks of manual abstraction now completes in hours. Analyze 6× more patients in the same timeframe compared to manual review workflows.
Improved Completeness
One standard OMOP CDM format for all data, >96% timeline completeness, confidence and audit trail, >99% de-identification accuracy.
Reuse at Scale
One standardized dataset supports cohorts, registries, analytics, and AI agents. Build the data foundation once, innovate endlessly on top. No redundant pipeline development, consistent results across applications.
Embedded Governance
Full provenance, lineage, and auditability of every derived clinical fact. Confidence scores for extracted data. Transparent handling of conflicts and missing data. Audit trails for regulatory compliance (HIPAA, GDPR).
Regulatory Readiness
Parallel identified and de-identified datasets kept semantically synchronized. Research models move to production without pipeline rewrites. Train on de-identified data, deploy on identified data, using identical feature definitions.
Future-Proof AI
A durable data foundation supporting advanced analytics, ML, and agentic systems at enterprise scale. New AI applications introduced without re-engineering data preparation. Evolves with advances in NLP and clinical AI.
The Result: A Repeatable Operating Model for Secondary Use of Clinical Data
The outcome is not just cleaner data, but a repeatable operating model for secondary use, where accuracy, trust, and reuse are built into the foundation rather than retrofitted for each new project.
Instead of spending 80% of effort on data wrangling for every new use case, teams spend 10% on data integration (once) and 90% on innovation (continuously). Instead of accepting 40% incompleteness and hoping it doesn't matter, organizations build on 96%+ complete patient representations and trust their results. Instead of isolated data pipelines that conflict and diverge, a single shared foundation ensures consistency across cohorts, registries, analytics, and AI applications.
💡 The Bottom Line
Secondary use of clinical data has enormous potential, but only if the data accurately represents what happened to patients. Patient Journey Intelligence provides the foundation: complete multimodal clinical data integration, healthcare-specific NLP, terminology normalization, clinical reasoning, longitudinal timelines, de-identification, provenance, and continuous updates.
Build the foundation once. Innovate on accurate, trustworthy, reusable patient data, forever.
FAQ
Secondary use means taking clinical data originally collected during patient care and repurposing it for research studies, quality improvement programs, population health analytics, disease registries, AI model development, public health surveillance, and healthcare operations optimization. Unlike primary use for direct treatment, secondary use works with historically accumulated data from multiple systems.
Unstructured EHR extraction is the process of automatically identifying and converting clinical information from free-text sources such as physician notes, discharge summaries, pathology reports, radiology narratives, and operative records, into structured, queryable data. Approximately 80% of healthcare data is unstructured. Without automated unstructured EHR extraction, analytics and AI systems can only access the minority of clinical information that clinicians entered into structured fields, systematically missing diagnoses, medications, social determinants, biomarkers, and clinical context that exist only in narrative text.
NLP medical entity extraction is the use of language models trained on clinical text to identify and classify medical entities such as diagnoses, medications, procedures, findings, anatomical locations, and their attributes, from unstructured clinical notes. Unlike general-purpose NLP, healthcare-specific models understand clinical nuance: negation ("no evidence of pneumonia"), assertion status (confirmed vs. ruled-out vs. family history), uncertainty, and temporal relationships. Patient Journey Intelligence uses NLP medical entity extraction to surface clinical facts that never appear in structured EHR fields, achieving 85–95% precision on clinical extraction tasks, approximately 30% more accurate than general-purpose LLMs.
The clinical data accuracy gap refers to the systematic inaccuracy that occurs when secondary use projects operate on incomplete patient data. Studies show that 40% of diagnoses, 81% of suicidality mentions, 68% of cancer staging data, and over 90% of social determinants of health exist only in unstructured clinical notes and are missed by structured-only approaches.
Structured EHR fields capture only a subset of clinically relevant information and frequently omit context such as certainty, negation, temporality, disease progression, adverse events, social factors, and clinician reasoning. Studies show that 87% of extracted clinical concepts exist solely in free-text narratives with no structured counterpart.
Patient Journey Intelligence addresses eight requirements: complete multimodal data integration, healthcare-specific NLP, terminology standardization (SNOMED CT, RxNorm, LOINC, ICD-10-CM), clinical reasoning and conflict resolution, longitudinal patient timelines, privacy and de-identification, provenance and auditability, and continuous updates with living datasets.
Healthcare-specific NLP models achieve 85–95% precision on clinical extraction tasks, approximately 30% more accurate than general-purpose LLMs. These models understand clinical context including negation, uncertainty, assertion status (confirmed vs. ruled-out vs. family history), and temporal relationships that general AI tools miss.
Patient Journey Intelligence uses healthcare-specific NLP that detects negation ("no evidence of pneumonia"), uncertainty, and assertion status in clinical text. This prevents common errors like treating a ruled-out condition as a confirmed diagnosis, which naive text search or general-purpose AI would miss.
Patient Journey Intelligence delivers 96%+ timeline completeness by extracting clinical facts from all data modalities: structured fields, clinical notes, scanned PDFs, imaging metadata, and claims data. This compares to 60% or less completeness with structured-only approaches that miss information in unstructured text.
Patient timeline construction that previously took weeks of manual abstraction now completes in hours with Patient Journey Intelligence. Organizations analyze 6× more patients in the same timeframe while achieving higher completeness and consistency than manual review workflows.
Patient Journey Intelligence is designed for clinical research teams, quality improvement departments, population health analysts, registry abstractors, data science groups, and healthcare IT leaders who need accurate, complete patient data for secondary use. The platform is most valuable when accuracy matters and structured-only data is insufficient.
Patient Journey Intelligence is not designed for primary clinical care delivery, real-time EHR documentation, or billing system replacement. Organizations that only need simple structured data extracts without NLP, terminology normalization, or multimodal integration may not require the platform's full capabilities.
Traditional clinical data warehouses store structured EHR extracts but miss 40%+ of clinical information in unstructured notes. Patient Journey Intelligence extracts facts from all modalities using healthcare-specific NLP, normalizes to standard vocabularies, resolves conflicts, maintains temporal relationships, and provides living datasets with full provenance—delivering complete secondary use rather than just data storage.
Patient Journey Intelligence provides complete lineage tracking from source document through extraction to final OMOP representation. Every clinical fact includes confidence scores, extraction method, and direct links to supporting evidence. This provenance chain supports HIPAA, 21 CFR Part 11, IRB requirements, and audit trails for research reproducibility.
Yes. Patient Journey Intelligence maintains parallel identified and de-identified datasets from the same source data, kept semantically synchronized with identical feature definitions. This enables seamless progression from research (de-identified) to production (identified) without pipeline rewrites. De-identification achieves 99%+ accuracy.
A living dataset is continuously updated as new clinical data arrives, rather than being a static snapshot that becomes stale. Patient Journey Intelligence keeps patient journeys current automatically, ensuring AI agents and analytics always operate on complete, up-to-date patient representations.