De-identified OMOP Research Registry
The Patient Journey Intelligence platform automatically maintains synchronized identified and de-identified OMOP datasets in parallel, enabling researchers to query compliant data instantly while operational systems use the same standardized patient journeys
Multimodal Clinical Data De-identification
This video demonstrates the interface visually without audio narration.
Clinical research depends on access to comprehensive patient data, but protecting patient privacy creates a fundamental bottleneck. The De-identified OMOP Research Registry eliminates this tradeoff by automatically maintaining parallel identified and de-identified versions of your OMOP patient journeys, synchronized in real-time, standardized to OMOP CDM, and ready for immediate querying. Researchers get instant access to compliant data without waiting for manual de-identification, while clinical operations use the same curated patient journeys with full PHI for care delivery and quality improvement.
The Research Data Access Bottleneck
Healthcare organizations collect rich clinical data during routine care, but making it available for research remains painfully slow and expensive.
The Gap: Manual De-Identification Can't Scale
Researchers need access to real-world clinical data to study treatment effectiveness, develop AI models, conduct epidemiological studies, and assess trial feasibility. But accessing this data requires removing Protected Health Information (PHI) to comply with HIPAA, GDPR, and institutional review board requirements. This creates persistent challenges:
-
4-8 Week De-Identification Delays Manual chart review and PHI removal take weeks or months per dataset request. By the time researchers receive data, clinical knowledge has advanced and patient populations have changed. Feasibility studies that could take hours stretch into months, delaying grant applications and trial recruitment.
-
Inconsistent De-Identification Methods Each research project re-invents de-identification pipelines. One team uses simple regex patterns and misses contextual PHI. Another applies over-aggressive masking that destroys clinical utility. Date-shifting approaches differ across studies, breaking temporal relationships. This inconsistency creates compliance risk and makes results non-reproducible.
-
Research-to-Production Gap Models trained on manually de-identified research exports fail when deployed on operational data because feature definitions differ, terminologies don't match, and preprocessing steps aren't documented. A readmission prediction model trained on research data can't run on live EHR feeds without extensive re-engineering.
-
Stale Snapshots Instead of Living Data Traditional research databases are point-in-time exports that quickly become outdated. Researchers work with 6-month-old data while recent clinical insights and emerging patient cohorts remain inaccessible. Continuous research questions require repeated manual extracts and endless IRB amendments.
-
Separate Infrastructure for Identified and De-Identified Data Organizations maintain duplicate databases, one identified for operations, one de-identified for research, doubling storage costs and creating version drift. Updates to operational data don't automatically flow to research databases, requiring manual synchronization and introducing inconsistencies.
Real-World Impact
Consider a health system developing an AI model to predict sepsis onset:
- Months waiting for de-identified training data as IT manually reviews clinical notes, redacts PHI, and exports database snapshots
- Model trained on 18-month-old data because that's the last approved research extract, missing recent protocol changes and emerging patient populations
- Feature definitions differ between research and production because research used manually processed notes while production uses live EHR extracts with different parsing logic
- Model fails at deployment because date references shifted differently, medication codes don't align, and lab values were normalized using inconsistent logic
- New cardiovascular research project starts from zero six months later, rebuilding similar de-identification pipelines despite identical underlying data needs
This cycle wastes resources, delays discoveries, and prevents organizations from compounding their AI and research investments across projects.
How Patient Journey Intelligence Eliminates the Research Data Bottleneck
Patient Journey Intelligence provides a fundamentally different approach: continuously synchronized identified and de-identified OMOP datasets that stay up-to-date in real-time, eliminating manual de-identification delays while ensuring research and operational systems use identical data foundations.
Parallel Datasets: One Pipeline, Two Compliant Outputs
The platform automatically maintains two synchronized versions of every patient journey, generated from the same source data pipeline:
Identified OMOP Dataset
Full PHI preserved for clinical operations, care coordination, quality improvement, and point-of-care AI applications
- Complete patient names, MRNs, addresses
- Exact dates and timestamps
- Provider identifiers and locations
- Contact information for care coordination
- Real-time updates as clinical data arrives
De-identified OMOP Dataset
HIPAA Safe Harbor and GDPR compliant with consistent pseudonyms, date-shifting, and PHI removal for research and external collaboration
- Pseudonymized patient IDs (consistent across visits)
- Date-shifted with temporal relationships preserved
- Geographic info limited to state/country level
- All 18 HIPAA identifiers removed or generalized
- Automatically synchronized with identified dataset
The Critical Advantage: Identical Semantics, Different Privacy Levels
Because both datasets are generated from the same automated pipeline, they maintain identical data semantics:
- Same feature definitions: A "diabetes diagnosis" means the same thing in both datasets, same extraction logic, same terminology mappings, same temporal windows
- Same data lineage: Both datasets trace back to the same source documents with identical transformation steps documented
- Same OMOP structure: Identical table schemas, concept IDs, vocabulary mappings, and relationship hierarchies
- Same update cadence: New clinical data flows into both datasets simultaneously, keeping them synchronized
This eliminates the research-to-production gap. Train an AI model on the de-identified research dataset, then deploy it directly on the identified operational dataset, no re-engineering, no feature drift, no surprises. The model sees the same data structure, terminologies, and temporal patterns in both environments.
Continuous Synchronization: Always-Current Research Data
Unlike traditional point-in-time research extracts, the de-identified OMOP dataset stays continuously synchronized with your clinical systems:
- New clinical documents arrive (notes, labs, imaging reports) from your EHR or data feeds
- AI-powered extraction processes unstructured content and maps findings to OMOP concepts
- Parallel updates occur simultaneously:
- Identified OMOP dataset updated with full PHI for operational use
- De-identified OMOP dataset updated with compliant pseudonyms and date-shifting
- Researchers query immediately without waiting for manual de-identification or IT requests
This means researchers always work with current patient populations, recent treatment patterns, and up-to-date outcomes, not stale snapshots from months ago.
The De-Identification Workflow: From Source Data to Compliant Research Datasets
Creating compliant, de-identified research datasets requires more than simply removing names and dates. The platform provides an end-to-end de-identification workflow that combines automated PHI detection, configurable anonymization rules, visual validation tools, and comprehensive audit trails, ensuring your research data meets regulatory standards while preserving clinical utility.
How De-Identification Works in Patient Journey Intelligence
The de-identification process operates across all your clinical data modalities (e.g. structured EHR fields, unstructured clinical notes, and medical imaging), using a centralized workflow that maintains consistency and provides full transparency.
Automated PHI Detection
Before any data can be safely shared for research, the platform must identify all Protected Health Information across diverse formats:
- Structured Data: Database fields containing patient names, MRNs, Social Security numbers, addresses, and contact information are automatically flagged based on field type and content patterns.
- Unstructured Clinical Text: Medical Language Models trained on clinical documentation analyze free-text notes, pathology reports, discharge summaries, and radiology reports to detect contextual PHI that simple pattern matching would miss, like "patient's employer is Johns Hopkins" or "lives on Elm Street near the fire station."
- Medical Imaging (DICOM): Both DICOM metadata headers (patient name, date of birth, study dates, device serial numbers) and burned-in text within the pixel data itself (like patient labels visible on X-rays or CT scans) are scanned for identifiers that must be removed or obscured.
Every piece of detected PHI is logged with its location, type, and confidence score, creating a transparent record of what was found and how it was handled.
Configurable Anonymization Profiles
Different research use cases have different privacy requirements. A multi-site federated study may require stricter de-identification than an internal quality improvement project. The platform uses anonymization profiles, a set of predefined rules that specify exactly how each type of PHI should be handled.
Pre-Built Profiles: The platform includes ready-to-use templates aligned with major regulatory frameworks:
- HIPAA Safe Harbor: Removes all 18 HIPAA identifiers with no exceptions
- GDPR Compliance: Addresses GDPR-specific sensitive attributes beyond HIPAA requirements
- Clinical Trial Redaction: Balanced approach retaining maximum clinical utility while meeting ICH GCP standards
- Institutional Custom Profiles: Organization-specific policies tailored to your IRB requirements
Profile Configuration: Each profile defines tag-level rules (drop the field, replace with a placeholder, apply cryptographic hashing), pixel-level detection thresholds for burned-in text, and pattern recognition settings for contextual PHI. Administrators can create custom profiles, clone and modify existing ones, or temporarily disable profiles for testing.
De-identification Profiles
This video demonstrates the interface visually without audio narration.
This flexibility ensures your de-identification approach matches your specific regulatory obligations and research needs without requiring code changes or IT involvement for each new project.
The De-Identification Dashboard
Researchers and data stewards manage all de-identification activities through a centralized dashboard that provides real-time visibility into processing status and compliance metrics.
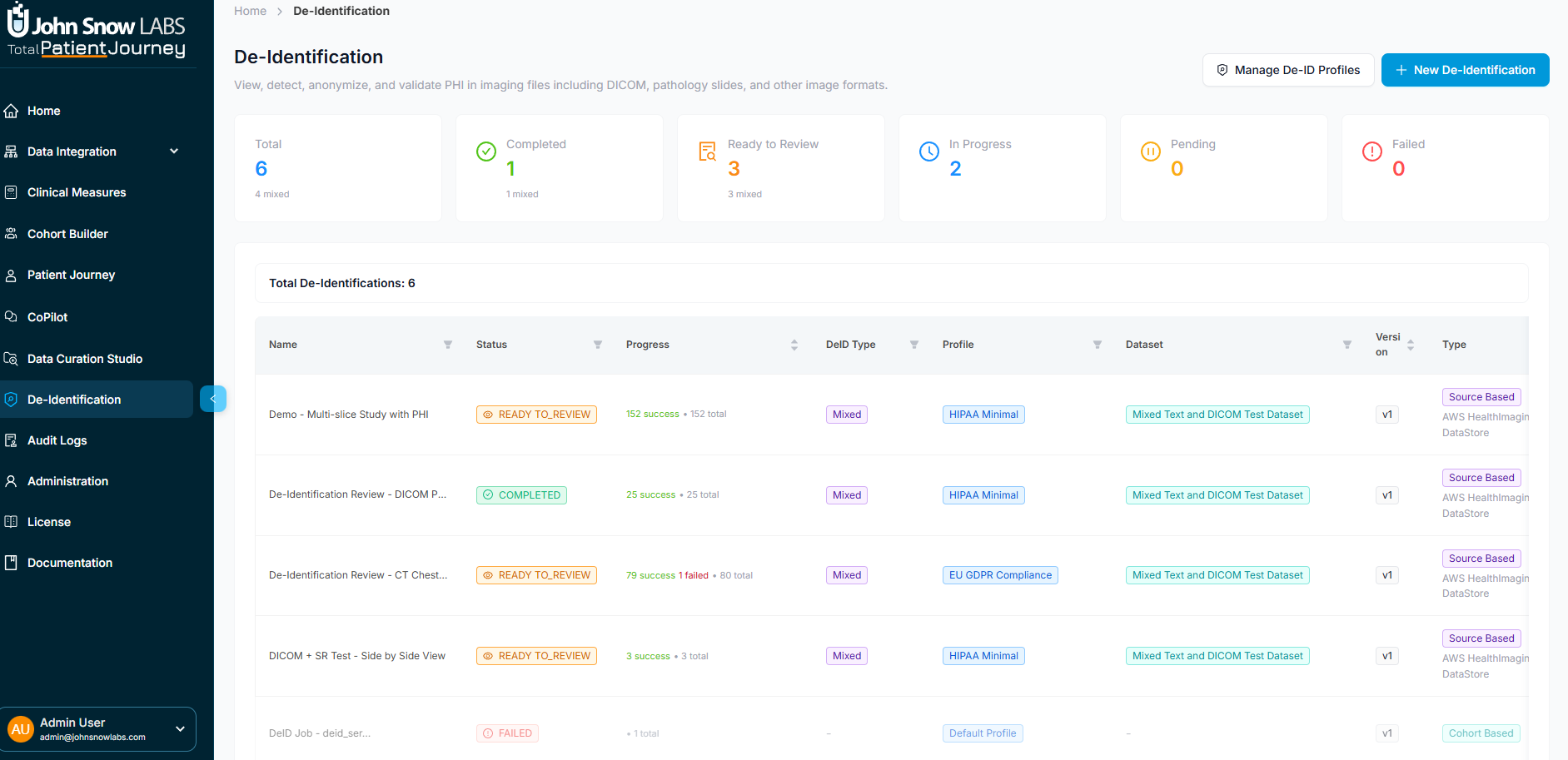
De-identification Dashboard showing job status, metrics, and management interface
System-Wide Metrics: At a glance, see total de-identification jobs executed, how many completed successfully, which are currently in progress, how many require manual review due to ambiguous PHI, and any that failed due to processing errors.
Job-Level Tracking: Each de-identification job displays its name, current status, which anonymization profile was applied, whether it processed a defined patient cohort or a broader data source, PHI detection statistics, processing timestamps, and available actions (view details, export results, edit the profile).
This transparency helps compliance officers demonstrate adherence to data governance policies and enables researchers to understand exactly what transformations were applied to their datasets.
Launching a De-Identification Job
Creating a new de-identified dataset is a guided process:
-
Select Your Data Source: Choose either a defined patient cohort (a specific group identified through your Data Curation workflows) or a broader data source like an external PACS system or cloud imaging repository.
-
Pick an Anonymization Profile: Select the regulatory template that matches your intended use, HIPAA Safe Harbor for external sharing, a lighter profile for internal-only research, or a custom profile defined by your institutional review board.
-
Initiate Processing: Confirm and launch. The system scans all patient records, clinical notes, lab results, imaging studies, and other data for PHI, applying your chosen anonymization rules automatically.
The platform processes data in parallel, handling large cohorts efficiently while maintaining detailed provenance for every transformation.
Visual Validation and Quality Assurance
Automated PHI detection is powerful, but clinical data is complex and nuanced. The platform provides human-in-the-loop validation tools to ensure de-identification quality before finalizing datasets for research use.
De-identification Visual Validation
This video demonstrates the interface visually without audio narration.
Patient Panel: A filterable sidebar lists all patients included in the de-identification job, showing each patient's age, sex, total clinical documents, and number of imaging studies. Selecting a patient loads their data for detailed review.
DICOM Viewer (Split View): For medical imaging, a dual-pane viewer displays the original image on the left and the de-identified version on the right, enabling precise visual comparison. Reviewers can scroll through slices, zoom and pan, adjust window/level settings, highlight detected PHI regions, and verify that burned-in patient labels were properly removed without damaging diagnostic content.
Metadata Inspector: A side-by-side comparison shows original DICOM headers next to de-identified versions, with a change summary highlighting which tags were modified, masked, or completely removed. This transparency ensures no critical clinical metadata was inadvertently lost during anonymization.
De-Identification Statistics Panel: For each imaging study, a structured summary reports:
- Total slices processed and how many contained detected PHI
- Pixel-based PHI detection results (detected instances, successfully cleared, rejected as false positives, pending manual review)
- Entity breakdown by PHI type (names, MRNs, institutions, dates, other identifiers)
- Metadata PHI enumeration showing which DICOM tags contained identifiers and their anonymization status
- Outcome summary (successful, failed, requires review)
- Pipeline metadata for auditing (profile used, execution time, processor ID, final status)
This level of detail supports both quality assurance and regulatory compliance documentation, providing IRB-ready evidence that PHI removal was thorough and systematic.
Export and Data Persistence
Once reviewed and validated, de-identified datasets can be used immediately or exported for external collaboration:
Export Locally: Generate DICOM files, CSV extracts, Parquet files, or FHIR bundles ready for submission to regulatory bodies, external research collaborators, or public data repositories.
Save to Internal Patient Journey Intelligence Storage: Persist the de-identified dataset within the platform for ongoing research access, enabling researchers to query it directly via SQL, REST API, or BI tools without repeated manual exports.
Link to OMOP CDM: Integrate de-identified imaging data with your broader OMOP Common Data Model, enabling combined analysis of imaging findings, lab results, medications, and clinical outcomes within a unified research database.
A final confirmation displays the number of exported studies, series, and slices, ensuring you know exactly what data was released.
Key De-Identification Capabilities
The de-identification workflow provides six integrated capabilities that work together to transform sensitive clinical data into compliant, research-ready datasets:
Automated PHI Detection Across All Modalities
Medical Language Models identify PHI in structured EHR fields, unstructured clinical notes, and DICOM imaging (both metadata headers and burned-in pixel content), catching contextual identifiers that pattern-matching tools miss.
Configurable, Standards-Aligned Anonymization Profiles
Pre-built templates for HIPAA Safe Harbor, GDPR, clinical trials, and institutional policies define exactly how each PHI type is handled, drop, replace, hash, or generalize, without requiring code changes for each research project.
Side-by-Side Visual Validation via DICOM Viewer
Dual-pane image viewer enables human reviewers to verify PHI removal quality by comparing original and de-identified versions slice-by-slice, ensuring burned-in text was removed without damaging diagnostic content.
Support for Cohort-Based and Source-Based Workflows
De-identify a specific patient cohort defined through Data Curation workflows, or process an entire data source like an external PACS system or cloud imaging repository, flexible workflows adapt to your research needs.
Detailed Audit and Processing Statistics
Comprehensive job-level metrics track PHI detection rates, anonymization outcomes, processing timestamps, and transformation lineage, providing IRB-ready documentation of compliance without manual chart audits.
Flexible Export Options for Research and Regulatory Workflows
Export de-identified data as local DICOM files, CSV/Parquet extracts, FHIR bundles, or save to internal Patient Journey Intelligence storage for direct querying, supporting both external collaboration and internal self-service research access.
This end-to-end workflow transforms de-identification from a months-long manual bottleneck into an automated, auditable process that maintains both regulatory compliance and clinical utility, enabling researchers to access compliant data in hours instead of waiting weeks for IT to manually redact PHI from point-in-time extracts.
De-Identification Compliance: HIPAA Safe Harbor and Expert Determination
The platform supports two de-identification pathways to meet different institutional needs and regulatory requirements.
HIPAA Safe Harbor Method
The Safe Harbor method provides straightforward compliance by removing or generalizing all 18 HIPAA identifiers. This approach requires no statistical analysis or external certification, if the specified identifiers are removed, the data is considered de-identified under HIPAA.
🔒 HIPAA Safe Harbor: 18 Required Removals
- Names (patient, relatives, employers)
- Geographic subdivisions smaller than state (addresses, cities, ZIP codes except first 3 digits)
- Dates directly related to the individual (birth, admission, discharge, death) - except year
- Telephone, fax, email
- Social Security numbers
- Medical record numbers
- Health plan beneficiary numbers
- Account numbers
- Certificate/license numbers
- Vehicle identifiers and serial numbers (license plates, VINs)
- Device identifiers and serial numbers (pacemakers, insulin pumps)
- Web URLs
- IP addresses
- Biometric identifiers (fingerprints, voiceprints, retinal scans)
- Full-face photos and comparable images
- Any other unique identifying number, characteristic, or code
Dates Handling: All dates are shifted by a consistent random offset per patient (e.g., +127 days), preserving temporal relationships between events while obscuring exact calendar dates. Ages over 89 are aggregated to "90+".
Consistent Obfuscation: Each patient receives the same obfuscation parameters across all records: the same date shift, the same pseudonym, ensuring referential integrity throughout the de-identified dataset. A patient's lab results, medications, visits, and procedures all maintain their relationships to the same de-identified patient ID, enabling longitudinal analysis while protecting privacy.
When to Use Safe Harbor:
- Straightforward compliance without statistical expertise required
- External data sharing where re-identification risk must be minimized
- Multi-site collaborations where consistent de-identification rules simplify governance
- Limited loss of data utility is acceptable (e.g., precise dates not needed for your analysis)
HIPAA Expert Determination Method
The Expert Determination method allows retention of more granular data (e.g., 5-digit ZIP codes, month-level dates, ages over 89) by having a qualified statistician certify that the re-identification risk is "very small." This preserves more clinical utility while maintaining HIPAA compliance.
📊 HIPAA Expert Determination: Risk-Based Assessment
Under the Expert Determination pathway, a qualified expert with statistical and scientific knowledge analyzes your dataset and certifies that the risk of re-identification is "very small." This allows retention of more detailed information than Safe Harbor while remaining HIPAA compliant.
What Can Be Retained:
- More precise geographic data (5-digit ZIP codes, county-level locations) if population density is sufficient
- Month and year of key clinical events (instead of year-only)
- Ages over 89 without aggregation if cohort size justifies it
- Rare diagnoses or procedures that might be indirectly identifying, if statistical risk assessment demonstrates low re-identification probability
John Snow Labs Expert Determination Service: Our team includes qualified statisticians who can perform the required risk assessment and provide formal Expert Determination documentation for your de-identified OMOP dataset. This service includes statistical disclosure control analysis, re-identification risk modeling, and comprehensive written certification that satisfies HIPAA requirements and institutional review board expectations.
When to Use Expert Determination:
- Preserving granular dates or geographic data is critical for your research questions
- Studying rare diseases or small populations where Safe Harbor might over-suppress data
- Internal use cases where additional utility justifies the expert review process
- You need formal documentation for IRB submissions or regulatory audits
GDPR Compliance: Beyond HIPAA Requirements
For organizations subject to the European Union's General Data Protection Regulation (GDPR), de-identification must address additional sensitive attributes beyond HIPAA's 18 identifiers. GDPR considers special categories of personal data that require explicit consent or lawful processing basis, and pseudonymization must account for these categories.
🔒 GDPR: Additional Sensitive Attributes
GDPR requires removal or pseudonymization of all personal data identifiers, including the core HIPAA identifiers plus additional sensitive attributes that could reveal:
Core Identifiers (overlaps with HIPAA):
- Names and surnames
- Postal addresses and precise location data
- Email addresses and phone numbers
- National identification numbers (passport, tax ID)
- Health insurance and medical record numbers
- Bank account and financial identifiers
- IP addresses and device identifiers
- Biometric data (fingerprints, facial images)
- Genetic data identifiers
GDPR-Specific Sensitive Attributes:
- Religious or philosophical beliefs (e.g., religious affiliation documented in social history)
- Trade union membership (documented in employment or insurance records)
- Political opinions or affiliations (e.g., mentioned in clinical notes)
- Sexual orientation or sexual life (documented in sexual history, partner gender references)
- Racial or ethnic origin (requires careful handling beyond clinical race/ethnicity categories)
- Criminal convictions or offenses (mentioned in social history, forensic evaluations)
GDPR requires additional documentation:
- Data Processing Agreements (DPA) with any third parties accessing the data
- Legitimate interest assessments or explicit consent documentation
- Data retention policies and automated deletion workflows
- Subject access rights mechanisms for patients to request their data or deletion
The platform's audit logging and provenance tracking support these GDPR requirements by documenting data lineage, access patterns, and transformation history.
Automated De-Identification Across All Data Modalities
Patient Journey Intelligence applies comprehensive de-identification across every data type in your clinical environment, ensuring consistent privacy protection regardless of where PHI appears.
Detect and Redact PHI from Unstructured Clinical Text
Medical Language Models trained on clinical documentation identify and remove PHI from free-text notes, discharge summaries, pathology reports, and radiology interpretations, handling medical jargon, abbreviations, and contextual references that generic NER tools miss.
Pseudonymize Structured EHR Fields
Database fields containing names, MRNs, SSNs, and identifiers are replaced with consistent pseudonyms across all tables. Each patient receives the same pseudonym throughout the de-identified dataset, preserving referential integrity for longitudinal analysis.
Shift Dates with Temporal Consistency
All dates for each patient are shifted by the same random offset, preserving time intervals between events (e.g., days between diagnosis and treatment start) while obscuring exact calendar dates. Ages over 89 are aggregated to "90+" per HIPAA requirements.
De-identify DICOM Imaging Metadata
DICOM headers containing patient names, birth dates, study dates, and device serial numbers are automatically scrubbed or pseudonymized, while preserving clinically relevant imaging metadata and study identifiers for cross-referencing.
Maintain Provenance and Audit Trails
Complete data lineage tracking documents which source records contributed to each de-identified patient journey, what transformations were applied, and when de-identification occurred, satisfying IRB and regulatory audit requirements without manual documentation.
Generate Validation Reports for Compliance
Automated reports summarize de-identification coverage (% of records processed), PHI detection rates, and compliance with Safe Harbor or Expert Determination requirements, providing IRB-ready documentation without manual chart audits.
Accessing Your De-Identified Research Data
The platform provides multiple access methods to fit your research workflows, from direct SQL queries to API integrations and batch exports.
Query Directly with SQL
Full PostgreSQL access to all OMOP CDM v5.4 tables with standard structure and vocabulary tables. Use your preferred SQL client or BI tools (DBeaver, DataGrip, Tableau) for complex cohort queries and longitudinal analysis.
Use for: Exploratory analysis, custom queries, BI dashboards
Integrate via REST API
Programmatic access with cohort definition endpoints, patient-level data retrieval, aggregate statistics, and FHIR R4-compatible resources. OAuth 2.0 authentication with role-based permissions.
Use for: Custom applications, automated pipelines, real-time integrations
Export for External Analysis
Full database dumps (PostgreSQL, Parquet), CSV/flat files for R/Python/SAS, FHIR bundles for interoperability, or custom formats (Spark, BigQuery, Snowflake) tailored to your analytical needs.
Use for: Statistical analysis, ML model training, external collaborations
What You Get: Measurable Outcomes from De-Identified Research Datasets
Organizations using the De-identified OMOP Research Registry report significant improvements in research productivity, compliance efficiency, and AI development velocity. By eliminating manual de-identification bottlenecks and maintaining synchronized datasets, institutions can compound their research investments across multiple use cases rather than rebuilding infrastructure for each project.
⚡ 95% Faster Time-to-Research
Eliminate 4-8 week de-identification bottlenecks. Researchers query pre-built de-identified data immediately, start analyzing in minutes instead of waiting months for manual PHI removal and data preparation.
🔄 Always-Current Research Data
Continuous synchronization means researchers always work with the latest clinical data. No stale snapshots, no manual refresh cycles, your research database stays current automatically as new patient data arrives.
🤖 Zero Feature Drift Between Research and Production
Train AI models on de-identified research data, deploy on identified operational data with identical semantics. Same feature definitions, terminologies, and temporal logic, eliminating the research-to-production gap that causes model failures.
🌐 Accelerate Multi-Site Studies
Standardized OMOP format enables seamless data sharing across institutions. Launch federated research networks without complex data harmonization or restrictive data use agreements.
🎯 10x More Research Output
Self-service access empowers researchers to explore multiple hypotheses and iterate quickly. Complete feasibility studies in hours, not weeks, dramatically increasing your institution's research productivity.
🏆 Competitive Grant Advantage
Demonstrate immediate data availability in grant applications. Provide preliminary data and feasibility evidence that reviewers demand, funded researchers can start analyzing on day one.
🔐 Enterprise-Grade Security
End-to-end encryption, role-based access controls, comprehensive audit logs, and flexible deployment options (cloud or on-premise). Air-gapped environments supported for maximum security requirements.
📈 Scale Research Without Scaling Staff
Automated workflows and self-service tools mean your existing team can support exponentially more research projects. Handle batch processing and high-volume queries without adding headcount.
🔍 Full Audit Trail & Reproducibility
Complete provenance tracking and version control ensure reproducible research. Every query, export, and analysis is logged, satisfying journal requirements and regulatory audits with zero manual effort.
Research Use Cases
The De-identified OMOP Research Registry supports a wide range of research applications, from retrospective outcomes studies to AI model development and multi-site collaborations. Because the data is already standardized to OMOP and continuously updated, researchers can launch new studies immediately without waiting for custom data preparation.
🔬 Retrospective Outcomes Studies
Analyze treatment effectiveness, disease progression, and long-term outcomes using real-world clinical data spanning multiple years of patient care.
Example: Comparative effectiveness of diabetes medications in elderly populations with comorbidities
🧪 Clinical Trial Feasibility
Assess patient availability and eligibility for proposed clinical trials before recruitment begins, reducing screen failures and accelerating enrollment timelines.
Example: Identify 200 treatment-naive metastatic breast cancer patients for Phase III trial in under 24 hours
📊 Comparative Effectiveness Research
Compare real-world outcomes across different treatment strategies, medications, or interventions using observational data that reflects actual clinical practice.
Example: Compare surgical vs. medical management of coronary artery disease in high-risk populations
🤖 AI Model Development
Generate curated training datasets with proper train/test splits, temporal validation sets, and feature engineering, all from standardized OMOP data that matches your production environment.
Example: Train predictive models for hospital readmission risk, then deploy on identified operational data with zero feature drift
🏥 Multi-Site Collaboration
Share standardized, de-identified datasets across institutions for federated research networks, rare disease studies, or comparative effectiveness analyses requiring larger sample sizes.
Example: Multi-center rare disease natural history studies with harmonized OMOP data from 10+ institutions
📈 Epidemiological Studies
Investigate disease incidence, prevalence, risk factors, and population trends using longitudinal patient journeys that capture complete care sequences over time.
Example: COVID-19 outcomes in immunocompromised populations, stratified by immunosuppression type and duration